A common problem at Pearl Harbor, biofouling affects harbors around the world. It’s the process by which barnacles, muscles, oysters and tubeworms accumulate on the bottom of boats and other surfaces. Now researchers at the University of Hawaiʻi at Mānoa’s Kewalo Marine Laboratory have discovered a biological trigger behind the buildup.
Crusty marine creatures begin life as miniscule larvae floating in the open ocean, says Michael Hadfield, a UH Mānoa professor of biology. But before the larvae settle on a surface and start to grow, they need a bacterial cue to initiate metamorphosis.
“The critical issue is how they find the right spot to make that transformation—the right place where food will be available and where there will be others of the species with which to reproduce,” Hadfield said. “The success of the species depends on the larvae settling in on exactly the right spot.”
Or the wrong spot to some—the U.S. Navy, commercial cargo shippers and many private boat owners would like to find a way to stop biofouling before it starts. A surface layer of barnacles or other marine life slows down boats and increases the amount of fuel it takes to move them through the water.
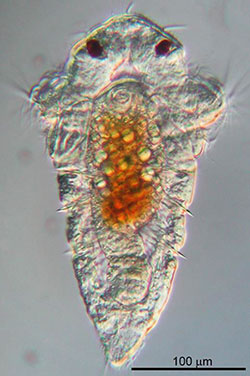
Biofouling begins when floating larvae come into contact with a biofilm formed by a microbe that coats steel, plastic and glass surfaces in calm ocean waters. New research from UH Mānoa and the California Institute of Technology has isolated the genetic underpinnings of this novel form of bacterium-animal interaction.
The results were published in the January 9, 2014, issue of the journal Science, in an article titled, “Marine tubeworm metamorphosis induced by arrays of bacterial phage tail-like structures.” Collaborator Nicholas Shikuma, the first author on this paper, studied with Hadfield and earned his masters degree at UH Mānoa. He is now a postdoctoral scholar at Caltech.
Hadfield and his team have been studying the marine tubeworm Hydroides elegans since 1990. In his laboratory, researchers cultivated strains of bacteria to identify the specific genes that are involved with causing the H. elegans larvae to abandon their free-swimming ways and “recruit” out of the ocean to begin a new life phase affixed onto harder surfaces.
Researchers found that these particular bacteria, Pseudoalteromonas luteoviolacea, produce arrays of phage tail-like structures that are similar to those produced for puncturing the cell membranes of competing bacteria. The phage tail-like structures appear to play a role in the bacterium-animal interaction as well, but researchers are still analyzing the specifics.
This bacteria’s not all bad. While a new layer of marine life on a previously pristine boat hull might be a negative for a boat owner, the same bacteria-driven process is a positive for repairing and restoring damaged reefs, for example. Greater knowledge of the forces that drive larval recruitment could also be a boon for oyster and clam growers in the mariculture industry, who rely on very similar processes to seed their stocks.
“Larval settlement is responsible for creating new communities on new surfaces,” Hadfield said. “But it’s also essential to continue those communities as old organisms die—to recruit new ones there to replace them. In the ecology of the sea, it’s one of the most important processes, and the more we understand about it, the better we can help it go.”
“Knowledge like this will help us to develop methods that target the process itself,” Hadfield said.
—By Talia Ogliore